AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Okazaki fragments dna ligase1/31/2024  The mitochondrial DNA of animal cells is a circular, double-stranded DNA that requires the activity of topoisomerase to be replicated. The mitochondria generate ATP as well as playing a role in programmed cell death and aging. Mitochondria Topoisomerase is also found in the mitochondria of cells. Read More: What is an example of didactic learning? Where is topoisomerase located? Responsible for patching together Okazaki fragments on the lagging strand during DNA replication. Which enzyme fixes the Okazaki fragments in the lagging strand?ĭNA Ligase DNA Ligase The enzyme responsible for sealing together breaks or nicks in a DNA strand. Because DNA ligase I is unable to join DNA to RNA, the RNA-DNA primers must be removed from each Okazaki fragment to complete lagging strand DNA synthesis and maintain genomic stability. Does DNA ligase remove primers?ĭNA ligase I is responsible for joining Okazaki fragments together to form a continuous lagging strand. This produces a series of disconnected Okazaki fragments. On the upper lagging strand, synthesis is discontinuous, since new RNA primers must be added as opening of the replication fork continues to expose new template. Why the lagging strand is synthesized discontinuously? … The leading strand is synthesized continuously while a lagging strand is synthesized in fragments which are called Okazaki fragments. What is the difference between and leading and lagging strand?Ī leading strand is the strand which is synthesized in the 5′-3’direction while a lagging strand is the strand which is synthesized in the 3′-5′ direction. A lagging strand requires a slight delay before undergoing replication, and it must undergo replication discontinuously in small fragments. Which is the lagging strand of DNA?Ī lagging strand is one of two strands of DNA found at the replication fork, or junction, in the double helix the other strand is called the leading strand. At the start of DNA replication, DNA unwinds and the two strands splits in two, forming two “prongs” which resemble a fork (thus, called replication fork). Relatively short fragment of DNA synthesized on the lagging strand during DNA replication. Okazaki fragments are short sequences of DNA nucleotides (approximately 150 to 200 base pairs long in eukaryotes) which are synthesized discontinuously and later linked together by the enzyme DNA ligase to create the lagging strand during DNA replication. Read More: What are the cells of cartilage? What are Okazaki fragments and where are they formed? 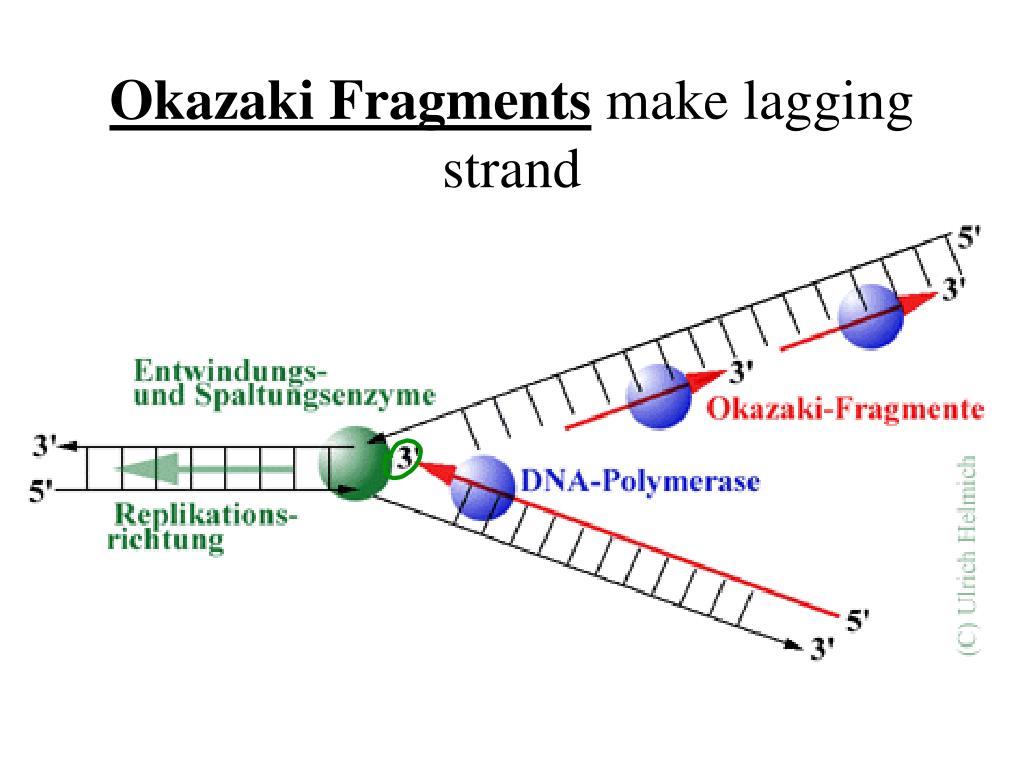
This lagging strand is synthesized in pieces because the DNA polymerase can only synthesize in the 5′ to 3′ direction, and so it constantly encounters the previously-synthesized new strand. 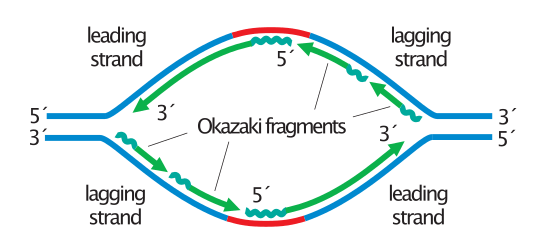
The “lagging strand” is synthesized in the direction away from the replication fork and away from the DNA helicase unwinds. Why is there a lagging strand in DNA replication? Why must there be a lagging strand during DNA synthesis? Explanation: The lagging strand exists because DNA is antiparallel and replication always occurs in the 5′ to 3′ direction. This leads to the formation of Okazaki fragments. As DNA polymerase can only add nucleotides in 5’→3′ direction of the growing strand, the lagging strand has to be synthesized discontinuously away from the replication fork. Okazaki fragments are necessary for the replication of both strands simultaneously. What are Okazaki fragments and why are they necessary? Polymerase I then removes RNA primers and fills the gaps between Okazaki fragments. Polymerase I In prokaryotic cells, polymerase III is the major replicative polymerase, functioning in the synthesis both of the leading strand of DNA and of Okazaki fragments by the extension of RNA primers. Which DNA polymerase removes Okazaki fragments? Okazaki fragments are short sequences synthesized in the lagging strand because DNA polymerase can synthesize only from 5′ to 3′, and the DNA strands are antiparallel. Which of the following statements best describes Okazaki fragments? They are formed in the lagging strand. Which best describes Okazaki fragments in DNA replication? On the lagging strand, DNA synthesis restarts many times as the helix unwinds, resulting in many short fragments called “Okazaki fragments.” DNA ligase joins the Okazaki fragments together into a single DNA molecule. On the leading strand, DNA synthesis occurs continuously. How does the Okazaki fragments occur for DNA replication? They are important because they allow for both daughter strands to be synthesized, which are necessary for cell division. 
Okazaki fragments are small sections of DNA that are formed during discontinuous synthesis of the lagging strand during DNA replication. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
